
LUO MIN
Assistant Professor
Department of Biological Sciences,
National University of Singapore,
14 Science Drive 4, Singapore 117543
66012310
dbslmin@nus.edu.sg
Academic Qualifications
Postdoc. (Harvard Medical School, USA), D. Phil. (U. of Missouri-Columbia, USA), B. Sc. (Shandong U., China)
Major Research Interests
Our laboratory is dedicated to unravelling the intricate processes of transmembrane translocation—a pivotal function in all living organisms. We employ a multidisciplinary structural biology approach, integrating cryo-electron microscopy (cryo-EM, Fig. 1) with an array of techniques from membrane reconstitution, biochemistry, molecular biology, biophysics, and cell biology. Our research primarily focuses on transmembrane proteins, such as ABC transporters and ATP synthase, which are critical to human health and disease. Furthermore, we innovate and implement cutting-edge methods to discover new membrane protein complexes within mitochondria
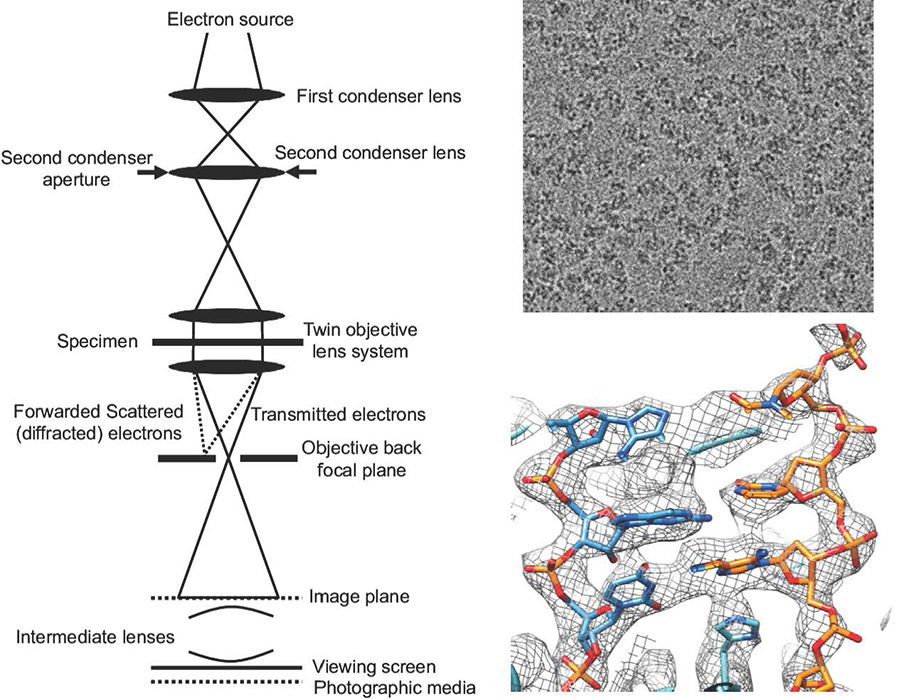
Figure 1. Single particle cryo-EM
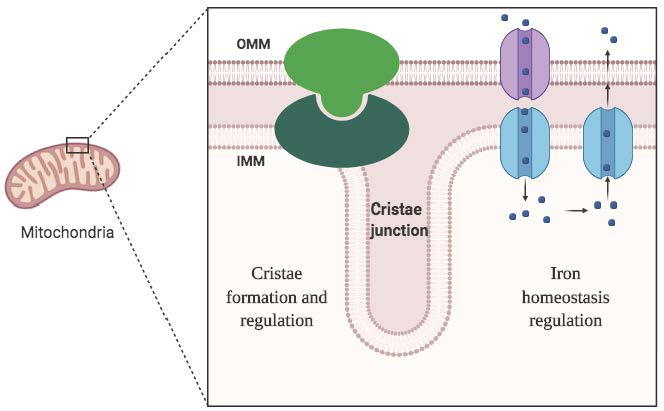
Figure 2. Proposed mitochondria study
Our current research on ABC transporters include two directions. First, we aim to elucidate the molecular underpinnings of cell division, utilizing prokaryotic systems as a model. Second, we delve into mitochondrial biology, concentrating on the regulation of energy and ion homeostasis (Fig. 2). Through the synergy of biochemical, cellular, and biophysical techniques, we strive to answer key questions surrounding vital drug targets associated with membrane proteins.
Selected publications since joining NUS
2024
-
Fan, W.; Luo, M.*. Structural View of Human ABCs in Multidrug Resistance. Biomolecules, 2024, (under revision). Preprints 2023, 2023121514. https://doi.org/10.20944/preprints202312.1514.v1
-
Jianwei Li #, Yutong He #, Xin Xu#, Martin Alcorlo, Jian Shi, David I. Roper, Juan A. Hermoso, Lok-To Sham, Luo M.*. Structural insights into peptidoglycan hydrolysis by the FtsEX system in Escherichia coli during cell division. eLife. 2024, (accepted, in press)
-
Conformational ensemble of Yeast ATP Synthase at Low pH Reveals Unique Intermediates and Plasticity in F1-Fo Coupling. Sharma, S*, Luo, M., Patel, H, Mueller, D. M* and Liao, M*. Nature Structure and Molecular Biology. 2024, (accepted, in press)
2023
-
Yu L.#, Xu X.#, Chua W.Z. #, Feng H.#, Ser Z., Shao K., Shi J., Wang Y., Li Z, Sobota R.M., Sham L.T.*, and Luo M.*. Structural basis of peptide secretion for Quorum sensing by ComA. Nature Communications. 2023, DOI: 10.1038/s41467-023-42852-9
-
Li J.#, Xu X.#, Shi J., Hermoso J.*, Sham L.T.*, and Luo M.*. Regulation of the Cell Division Hydrolase RipC by the FtsEX system in Mycobacterium tuberculosis. Nature Communications. 2023, DOI: 10.1038/s41467-023-43770-6
-
Xu, X.#; Li, J.#; Chua, W. Z.; Pages, M. A.; Shi, J.; Hermoso, J. A.*; Bernhardt, T.*; Sham, L. T.*; Luo, M.*. Mechanistic insights into the regulation of cell wall hydrolysis by FtsEX and EnvC at the bacterial division site. Proc Natl Acad Sci USA. 2023, 120 (21), e2301897120.
-
Yeow J, Luo M., and Chng S.S.. Molecular mechanism of phospholipid transport at the bacterial outer membrane interface. Nature Communications. 2023, 14(1):8285. DOI: 10.1038/s41467-023-44144-8.
-
Dutta C, Krishnamurthy P, Su D, Sung H, Collie G, Pasco M, Marzinek J, Bond P, Verma C, Grélard A, Loquet A, Li J, Luo M., Barboiu M*, Guichard G, R*. Kini M*, Kumar P*. Nature-inspired synthetic oligourea foldamer channels allow water transport with high salt rejection. CHEM. 2023, DOI: https://doi.org/10.1016/j.chempr.2023.04.007
-
Zeng, T.; Yin, J; Liu, Z; Li, Z; Zhang, Y; Lv, Y; Lu, M; Luo, M.; Chen, M; Xiao, Y. Mechanistic insights into transposon cleavage and integration by TnsB of ShCAST system. Cell Reports. 2023, Jul 25;42(7):112698.
-
Zhang Y*, Ding C, Chen Y, Wu M, Luo M., Editorial: Fetal phenotypes of rare diseases: application and evaluation of prenatal exome sequencing and pathogenesis research of rare diseases. Front Genet. 2023, 14, 1205726
Book chapter:
- Mishra K., Luo M.*. Mitochondrial Channels and Their Role in Cardioprotection. Book chapter. Ion Transporters – From Basic Properties to Medical Treatment. 2021
